pyinaturalist¶



Python client for the iNaturalist APIs. See full documentation at https://pyinaturalist.readthedocs.io.
Installation¶
Install with pip:
$ pip install pyinaturalist
Or, if you would like to use the latest development (non-stable) version:
$ pip install --pre pyinaturalist
Or, to set up for local development (preferably in a new virtualenv):
$ git clone https://github.com/niconoe/pyinaturalist.git
$ cd pyinaturalist
$ pip install -Ue ".[dev]"
Development Status¶
Pyinaturalist is under active development. Currently, a handful of the most relevant API endpoints are implemented, including:
Searching, creating, and updating observations and observation fields
Searching for species
See below for some examples, see Reference for a complete list, and see Issues for planned & proposed features. More endpoints will continue to be added as they are needed. PRs are welcome!
Note
The two iNaturalist APIs expose a combined total of 103 endpoints. Many of these are primarily for internal use by the iNaturalist web application and mobile apps, and are unlikely to be added unless there are specific use cases for them.
Examples¶
Observations¶
Search all observations matching a criteria:¶
from pyinaturalist.node_api import get_all_observations
obs = get_all_observations(params={'user_id': 'niconoe'})
See available parameters.
For authenticated API calls, you first need to obtain a token for the user:¶
from pyinaturalist.rest_api import get_access_token
token = get_access_token(username='<your_inaturalist_username>', password='<your_inaturalist_password>',
app_id='<your_inaturalist_app_id>',
app_secret=<your_inaturalist_app_secret>)
Note: you’ll need to create an iNaturalist app.
Create a new observation:¶
from pyinaturalist.rest_api import create_observations
params = {'observation':
{'taxon_id': 54327, # Vespa Crabro
'observed_on_string': datetime.datetime.now().isoformat(),
'time_zone': 'Brussels',
'description': 'This is a free text comment for the observation',
'tag_list': 'wasp, Belgium',
'latitude': 50.647143,
'longitude': 4.360216,
'positional_accuracy': 50, # meters,
# sets vespawatch_id (an observation field whose ID is 9613) to the value '100'.
'observation_field_values_attributes':
[{'observation_field_id': 9613,'value': 100}],
},
}
r = create_observations(params=params, access_token=token)
new_observation_id = r[0]['id']
Upload a picture for this observation:¶
from pyinaturalist.rest_api import add_photo_to_observation
r = add_photo_to_observation(observation_id=new_observation_id,
file_object=open('/Users/nicolasnoe/vespa.jpg', 'rb'),
access_token=token)
Update an existing observation of yours:¶
from pyinaturalist.rest_api import update_observation
p = {'ignore_photos': 1, # Otherwise existing pictures will be deleted
'observation': {'description': 'updated description !'}}
r = update_observation(observation_id=17932425, params=p, access_token=token)
Get a list of all (globally available) observation fields:¶
from pyinaturalist.rest_api import get_all_observation_fields
r = get_all_observation_fields(search_query="DNA")
Sets an observation field value to an existing observation:¶
from pyinaturalist.rest_api import put_observation_field_values
put_observation_field_values(observation_id=7345179,
observation_field_id=9613,
value=250,
access_token=token)
Get observation data in alternative formats:¶
A separate endpoint can provide other data formats, including Darwin Core, KML, and CSV:
from pyinaturalist.rest_api import get_observations
obs = get_observations(user_id='niconoe', response_format='dwc')
Taxonomy¶
Search for all taxa matching some criteria:¶
Let’s say you partially remember either a genus or family name that started with ‘vespi’-something:
>>> from pyinaturalist.node_api import get_taxa
>>> response = get_taxa(q="vespi", rank=["genus", "family"])
>>> print({taxon["id"]: taxon["name"] for taxon in response["results"]})
{52747: "Vespidae", 84737: "Vespina", 92786: "Vespicula", 646195: "Vespiodes", ...}
Oh, that’s right, it was ‘Vespidae’! Now let’s find all of its subfamilies using its taxon ID from the results above:
>>> response = get_taxa(parent_id=52747)
>>> print({taxon["id"]: taxon["name"] for taxon in response["results"]})
{343248: "Polistinae", 84738: "Vespinae", 119344: "Eumeninae", 121511: "Masarinae", ...}
Get a specific taxon by ID:¶
Let’s find out more about this ‘Polistinae’ genus. We could search for it by name or by ID, but since we already know the ID from the previous search, let’s use that:
>>> from pyinaturalist.node_api import get_taxa_by_id
>>> response = get_taxa_by_id(343248)
There is a lot of info in there, but let’s just get the basics for now:
>>> basic_fields = ["preferred_common_name", "observations_count", "wikipedia_url", "wikipedia_summary"]
>>> print({f: response["results"][0][f] for f in basic_fields})
{
"preferred_common_name": "Paper Wasps",
"observations_count": 69728,
"wikipedia_url": "http://en.wikipedia.org/wiki/Polistinae",
"wikipedia_summary": "The Polistinae are eusocial wasps closely related to the more familiar yellow jackets...",
}
Taxon autocomplete¶
This is a text search-optimized endpoint that provides autocompletion in the Naturalist web UI:
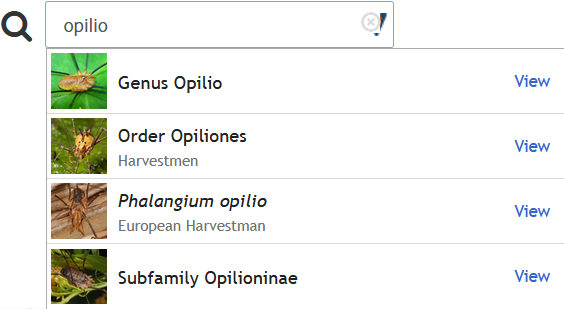
This one is a bit more niche, but it provides a fast way to search the iNaturalist taxonomy database. Here is an example that will run searches from console input:
from pyinaturalist.node_api import get_taxa_autocomplete
while True:
query = input("> ")
response = get_taxa_autocomplete(q=query, minify=True)
print("\n".join(response["results"]))
Example usage:
> opilio
527573: Genus Opilio
47367: Order Opiliones (Harvestmen)
84644: Species Phalangium opilio (European Harvestman)
527419: Subfamily Opilioninae
...
> coleo
372759: Subclass Coleoidea (Coleoids)
47208: Order Coleoptera (Beetles)
359229: Species Coleotechnites florae (Coleotechnites Flower Moth)
53502: Genus Brickellia (brickellbushes)
...
<Ctrl-C>
If you get unexpected matches, the search likely matched a synonym, either in the form of a
common name or an alternative classification. Check the matched_term property for more
info. For example:
>>> first_result = get_taxa_autocomplete(q='zygoca')['results'][0] >>> first_result["name"] "Schlumbergera truncata" >>> first_result["matched_term"] "Zygocactus truncatus" # An older synonym for Schlumbergera
Dry-run mode¶
While developing & testing an application that uses an API client like pyinaturalist, it can be useful to temporarily mock out HTTP requests, especially requests that add, modify, or delete real data. Pyinaturalist has some settings to make this easier.
Dry-run all requests¶
To enable dry-run mode, set the DRY_RUN_ENABLED variable. When set, requests will not be sent
but will be logged instead:
>>> import logging
>>> import pyinaturalist
# Enable at least INFO-level logging
>>> logging.basicConfig(level='INFO')
>>> pyinaturalist.DRY_RUN_ENABLED = True
>>> get_taxa(q='warbler', locale=1)
{'results': [], 'total_results': 0}
INFO:pyinaturalist.api_requests:Request: GET, https://api.inaturalist.org/v1/taxa,
params={'q': 'warbler', 'locale': 1},
headers={'Accept': 'application/json', 'User-Agent': 'Pyinaturalist/0.9.1'}
Or, if you are running your application in a command-line environment, you can set this as an environment variable instead (case-insensitive):
$ export DRY_RUN_ENABLED=true
$ python my_script.py
Dry-run only write requests¶
If you would like to run GET requests but mock out any requests that modify data
(POST, PUT, DELETE, etc.), you can use the DRY_RUN_WRITE_ONLY variable
instead:
>>> pyinaturalist.DRY_RUN_WRITE_ONLY = True
# Also works as an environment variable
>>> import os
>>> os.environ["DRY_RUN_WRITE_ONLY"] = 'True'
If you have any suggestions or questions about pyinaturalist feel free to email me at nicolas@niconoe.eu.
If you encounter any errors or problems with pyinaturalist, please let me know! Open an Issue at the GitHub http://github.com/niconoe/pyinaturalist main repository.